step-onset (run-onset.xml)
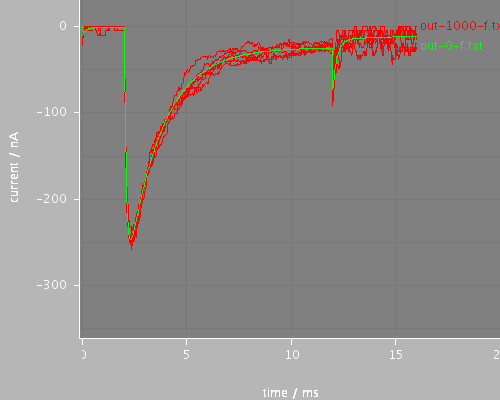
Voltage clamp step onset behavior for continuous and stochastic channels. The step begins at 1.985ms and the timestep is 0.02ms. This uses the midpoint evaluation method for the step, (stepStyle = midpoint in the voltage clamp) so the effect is as if the step started at 1.980ms. This fugure should be compared with the file run-onset-avg, which does the same calculation but with stepStyle = average for the clamp.
Total CPU time 0.2210 seconds; at 16:38:53 Fri 28 Sep 2007
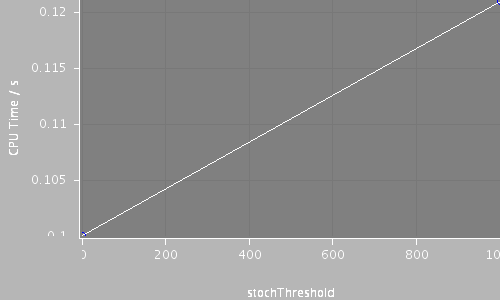
| Compartments | Stochastic channels / cpmts | Continuous channels / cpmts | time/ms | timestep/ms | stochThreshold | CPU Time / s |
| 1 | 0 / 0 | 201 / 1 | 16.0 | 0.02 | 0 | 0.1 |
| 1 | 201 / 1 | 0 / 0 | 16.0 | 0.02 | 1000 | 0.121 |
 |
Predefined views
whole
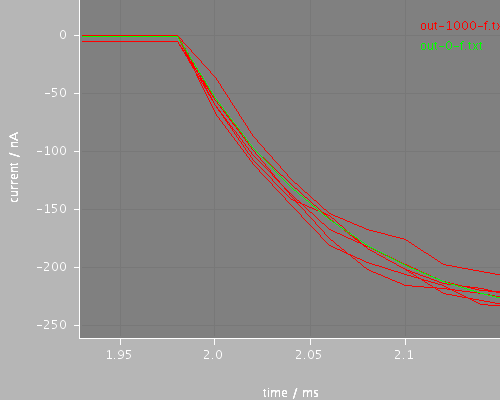
start
All files
| Model | Preprocessed | Outupt data | Reference data etc |
| run-onset.xml cell.xml membrane.xml environment.xml recording-short.xml na1.xml summary.xml view.xml recording-short-avg.xml |
out-16-fdata.txt out-0-fdata.txt out-1000-fdata.txt |
out-16-f.dat out-16-f.txt out-0-f.dat out-0-f.txt out-1000-f.dat out-1000-f.txt |
step-onset.jar |
Model
Archive file of the complete model: step-onset.jarrun-onset.xml
<PSICSRun timeStep="0.02ms" runTime="16ms" startPotential="-65mV"
morphology="cell"
environment="environment"
properties="membrane"
access="recording-short"
stochThreshold="0"
repeats="20">
<StructureDiscretization baseElementSize="3um"/>
<RunSet vary="stochThreshold" values="[0, 1000]"
filepattern="out-$.txt">
</RunSet>
<About>
Voltage clamp step onset behavior for continuous and stochastic channels. The step begins
at 1.985ms and the timestep is 0.02ms. This uses the midpoint evaluation method for the step,
(stepStyle="midpoint" in the voltage clamp)
so the effect is as if the step started at 1.980ms. This fugure should be compared with
the file run-onset-avg, which does the same calculation but with stepStyle="average" for the clamp.
</About>
<ViewConfig>
<LineGraph width="500" height="400">
<XAxis min="0" max="250" label="time / ms"/>
<YAxis min="-80" max="60" label="current / nA"/>
<LineSet file="out-1000-f.txt" color="red" maxshow="6"/>
<LineSet file="out-0-f.txt" color="green" maxshow="2"/>
<View id="whole" xmin="0." xmax="20." ymin="-360." ymax="30."/>
<View id="start" xmin="1.93" xmax="2.15" ymin="-260." ymax="30."/>
</LineGraph>
</ViewConfig>
</PSICSRun>
cell.xml
<CellMorphology id="cell">
<Point id="p0" x="0" y="0" z="0" r="1."/>
</CellMorphology>
membrane.xml
<CellProperties id="membrane"
cytoplasmResistivity="100ohm_cm"
membraneCapacitance="1uF_per_cm2">
<ChannelPopulation id="napop" channel="na1" density="16per_um2"/>
</CellProperties>
environment.xml
<CellEnvironment id="environment"> <Ion id="Na" name="Sodium" reversalPotential="80mV"/> <Ion id="K" name="Potassium" reversalPotential="-80mV"/> </CellEnvironment>
recording-short.xml
<Access id="recording-short" saveInterval="0.05ms"> <VoltageClamp at="p0" lineColor="red" hold="-80.0mV" stepValue="midpoint"> <VoltagePulse id="vstep" start="1.985ms" to="30mV" duration="10ms"/> </VoltageClamp> </Access>
na1.xml
<KSChannel id="na1" gSingle="30pS" permeantIon="Na"> <ClosedState id="c1"/> <OpenState id="o1"/> <ClosedState id="c2"/> <VHalfTransition from="c1" to="o1" vHalf = "-35mV" z="4.5e" gamma="0.8" tau="0.55ms" tauMin="0.1ms"/> <VHalfTransition from="o1" to="c2" vHalf = "-70mV" z="1.1e" gamma="0.90" tau="12.0ms" tauMin="1.5ms"/> </KSChannel>
recording-short-avg.xml
<Access id="recording-short" saveInterval="0.05ms"> <VoltageClamp at="p0" lineColor="red" hold="-80.0mV" stepValue="average"> <VoltagePulse id="vstep" start="1.985ms" to="30mV" duration="10ms"/> </VoltageClamp> </Access>